More and more data in healthcare has been transferred to a digital format over the past decade. Furthermore, latest advances in hardware enabled the use of Artificial Intelligence (AI) algorithms for more and more use cases, also in digital health. On the one hand, the use of real-world healthcare data is expected to have great potentials for building highly accurate AI-based decision support tools, such as Clinical Prediction Models (CPMs). On the other hand, healthcare data has to be considered as very sensitive data, which requires additional protection measures. For example, the European General Data Protection Regulation (GDPR) and the Canadian Personal Information Protection and Electronic Documents Act (PIPEDA) highlight Personal Health Information (PHI) in their regulation as dedicated type of data.
Thanks to the contributing experts from our partners for their input for this article: Dr. Oliver Günther (UBC), Konstantin Pandl (KIT), Aadil Rasheed (HPI), and Dr. Matthieu-P. Schapranow (HPI). Photo by Marvin Meyer.
Federated Learning: A privacy-preserving way for use of AI in healthcare
Real-world data builds the foundation for defining CPMs, e.g., to predict the risk for graft failure after transplantation. The quality of such CPMs depends on the variability of incorporated real-world data to identify connection between features and clinical outcomes. For example, the data from one transplant center might differ from another center due to patient population, gender distribution, presence of patient comorbidities, etc.
The NephroCAGE consortium builds on latest research results on Federated Learning Infrastructures (FLIs) to bridge this gap. FLIs enable the transfer of AI algorithms to data, which resides locally in their protected sites. Such a decentralized FLI system opens the door for using AI on privacy-focused use cases, such as kidney transplantation in the NephroCAGE consortium.
How to design clinical predictive models for kidney disease
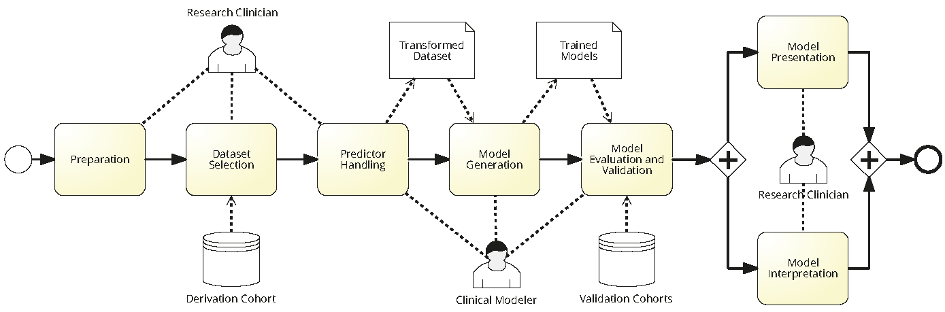
Designing CPMs requires Clinicians and Modeler to work collaboratively together. Figure 1 depicts a BPMN model showing various steps in modeling a CPM with all the actors involved in the process along with generated artifacts. First, a research clinician prepares requirements and selects relevant datasets from the derivation cohort. Later, a clinical modeler uses the selected dataset for training CPMs. After completing the training, the modeler validates the CPMs on a special dataset, the validation cohort. Finally, the research clinician assess the quality and results of the CPMs with regards to their clinical applicability.
How to incorporate patient data for clinical predictive models?
The use of real-word clinical data is crucial for the prediction quality of the CPMs. There are multiple ways to incorporate real-world clinical data in privacy-compliant ways. In the following, selected alternatives are outlined.
- Acquiring informed consent from prospective patients. Typically, consent management is covered as an integral aspect of the clinical treatment contract. Therefore, patients will receive a lay-understandable explanation of how and for what purpose their data will be used. Every patient can decide individually to share their data for the purpose of developing CPMs.
- A very innovative approach to obtain patient consent even after leaving the hospital is the use of data donation. Citizens will be offered digital tools to manage their personal consent to share personal healthcare data for specific purposes, e.g., donate de-identified data of my kidney transplant for developing a CPM. Such digital tools offer a digital communication channel to highly motivated patients interested to improve healthcare.
- Data de-identification for retrospective data. This option is helpful if there is no easy-to-adopt option to obtain an explicit patient consent, e.g., because the data was acquired in the past and a reach out to the patient failed. One option is to remove personal patient identifiers from the data, e.g., name, birthday, address. Although this is a complex task due to the variety of data types and identifiers, there is limited tool support available for that purpose, e.g. tools of the MOSAIC project help to evaluate the risk of reidentification in the de-identified dataset.
A central repository of de-identified patient data must be considered as sensitive data upon the nature of data. It still requires specific measures to ensure privacy and data security. Kidney transplant dataset possess elevated risk because the number of annually performed kidney transplantation is low compared to other surgeries. Furthermore, requested lab values and clinical interventions ordered for kidney transplant patients need to be considered as very specific compared to the remaining patient population. Moreover, forming a multi-site database of transplant data requires extensive technical work and legal knowledge. If the database spans multiple regions or even countries additional national law needs to be respected.
Federated learning and how it can facilitate the use of real-world clinical data for AI
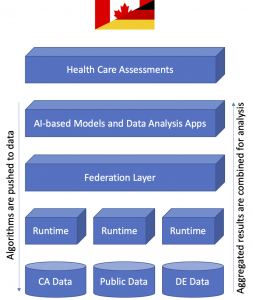
Fig. 2: Software architecture for federated learning. Data resides in their decentralized silos and algorithms are exchanged via the federation layer.
Federated learning is considered as paradigm shift for development of AI models. Instead of aggregating sensitive data from distributed clinical sources, algorithms are transferred to where the data resides. Thus, the processing of clinical data can occur distributed and in parallel without the need to send sensitive data around. As a result, sensitive clinical data stays in its original location and does not leave hospitals. Instead, only aggregated results of the data analysis are shared in compliance with data protection regulations. The FLI can be seen as an additional layer on top of the clinical IT infrastructure, which enables the exchange of models and model-specific parameters between participating sites as depicted in Figure 2.
FLIs can be implemented following different architecture principles, for example, either as a centralized or decentralized infrastructure. On the one hand, centralized FLIs come with advantages in service management, but might become a single point of failure in case of outages. On the other hand, decentralized FLIs are more robust against service attacks, but require additional management overhead. For the NephroCAGE project, we prefer the use of a decentralized infrastructure because individual clinical sites can be added to the mash-up and CPMs can be exchanged between parties on individual agreements. Furthermore, the decentralized FLI is more robust and less prone to suffer from complete outages.
Summary
In our NephroCAGE consortium, we incorporate federated learning infrastructure for the application of AI with healthcare data. It provides the foundation for the sharing of AI algorithms. Hence, sensitive transplant data remains at local clinical sites, while algorithms are transferred to clinical sites for data processing.




No responses yet